Biography
Yu Zhu is now a MSc Drug Design student at University College London, United Kingdom and he was an undergraduate student majoring in Bioinformatics at Soochow University, China during the period of 2018 and 2022.
Under the guidance of Assoc. Prof. Guang Hu and Assoc. Prof. Zhongjie Liang, he is now focusing on how to integrate bioinformatics and biophysics methods to have an insight into protein dynamics, along with protein sequence, structure and function research and finally figure out how to use structural information for the construction and analysis of protein-protein interaction networks, paving the way for drug discovery. Also, supervised by Prof. Bairong Shen at Institutes for Systems Genetics, West China Hospital, he established a risk factor knowledgebase for neurodegenerative diseases (NDDRF) together with other group members.
This personal page will no longer be updated.
Check Personal CV here.
- Computational Biology
- Systems Biology
- Structure Bioinformatics
- Protein Sequence & Structure & Dynamics & Function
- Biological Network
- Bioinformatics Algorithm
- Proteome Informatics
- Allosteric Drug Design
- Database
-
BSc Bioinformatics, 2018 - 2022
Soochow University, China
-
MSc Drug Design, 2022 - Present
University College London, United Kingdom
Skills
Commonly used programming language
Commonly used programming language
Commonly used operating system
Establishment of biological database and web page
Multi-scale bioinformatics analysis
Algorithms related to biological data processing, network topology, machine learning …
Research Experiences
Supervised by: Assoc. Prof. Guang Hu and Assoc. Prof. Zhongjie Liang
Responsibilities include:
- Construct the biological network of the complex, and develope indicators that quantitatively characterize the characteristics of the mutant residues
- Predict the allosteric site and allosteric path of the complex based on perturbation response scanning (PRS) analysis
- Explore pocket information in the trajectory of DNMTs, evaluate their targetability, and determine potential allosteric sites combining mutation information
Supervised by: Assoc. Prof. Guang Hu and Assoc. Prof. Zhongjie Liang
Responsibilities include:
- Obtain the NSCLC-related disease network module from the pre-omics analysis
- Carry out research in five aspects, including functional enrichment analysis, structural modeling of protein interactions, mapping of mutation and post-translational modification information and druggability based on drug pockets
- Verify the pathogenic function of the NSCLC network module at the two levels of pathway and structure
Supervised by: Prof. Bairong Shen
Responsibilities include:
- Sort out the subject development process of glycomics, glycobiology, and glycobioinformatics, and learned about the forefront progress of the subject.
- Participate in several summit forums discussing frontier science in bioinformatics
Supervised by: Prof. Bairong Shen
Responsibilities include:
- Learn about the application of bioinformatics-related tools and database, and the data processing method of multi-omics analysis
- Join Prof. Shen Bairong’s research group, and take charge of data mining and cleaning for more than ten neurodegenerative diseases
- Establish the risk factors database for neurodegenerative diseases (NDDRF) together with other group members
- Follow up with the latest research progress in bioinformatics, and participate in academic conferences
Honors and Awards
- Outstanding Graduate of Soochow University 2022
- Outstanding Undergraduate Design (Thesis) of Soochow University 2022
- National Software Copyright (Certificate registration number: 2022SR0518920)
- National Software Copyright (Certificate registration number: 2022SR0518921)
- The First Prize for China Undergraduate Life Science Contest
- The Second Prize in the First “Zhaoyin” Cup Financial Mathematical Modeling Competition
- The Third Prize for Band C in 2021 National English Competition for College Students
- The Second Prize in the 22nd “Challenge Cup” Medical College of Soochow University Undergraduate Curricular Academic Science and Technology Works Competition
- National Scholarship
- Comprehensive Scholarship
- Merit Student
- The Special Scholarship for Innovation and Entrepreneurship
- The Special Prize Scholarship for Academic Excellence
- The First Prize Scholarship for Academic Excellence
- Comprehensive Scholarship
- The Special Scholarship for Innovation and Entrepreneurship
- Merit Student
- Scientific and Technological Innovation Star of Medical College
- The First Prize for Academic Excellence
- Comprehensive Prize
- Merit Student
- The Special Prize for Spiritual Civilization
- Excellent Volunteer Team
- Excellent Summer Social Practice Team
Posts
Publications
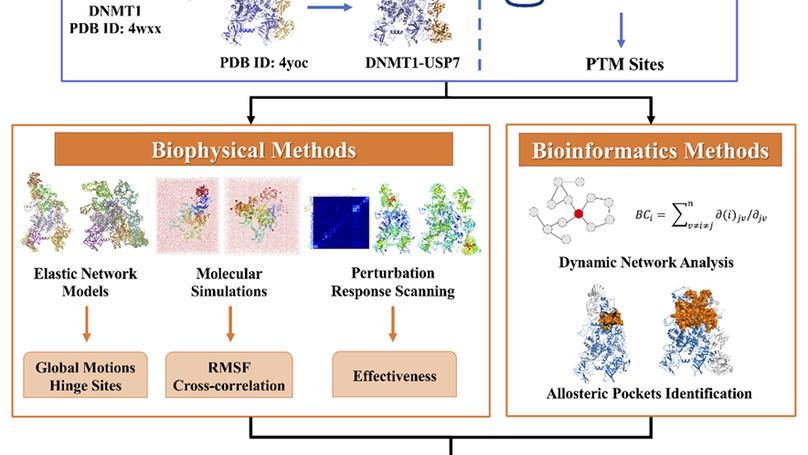
Allosteric drug design targeting DNMT1 Protein-Protein Interactions (PPIs) combining both biophysics and bioinformatics methods.
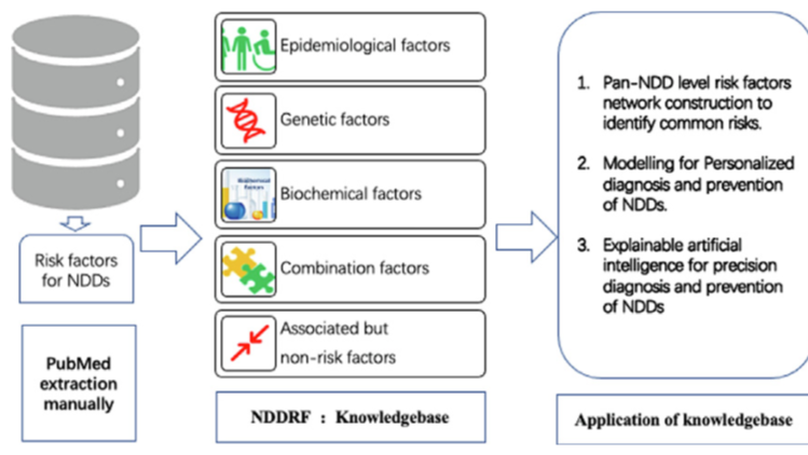
NDDRF provides the structured information and knowledge resource on risk factors of NDDs. It could benefit the future systematic and personalized investigation of pan-NDDs genesis and progression. Meanwhile it may be used for the future explainable artificial intelligence modeling for smart diagnosis and prevention of NDDs.
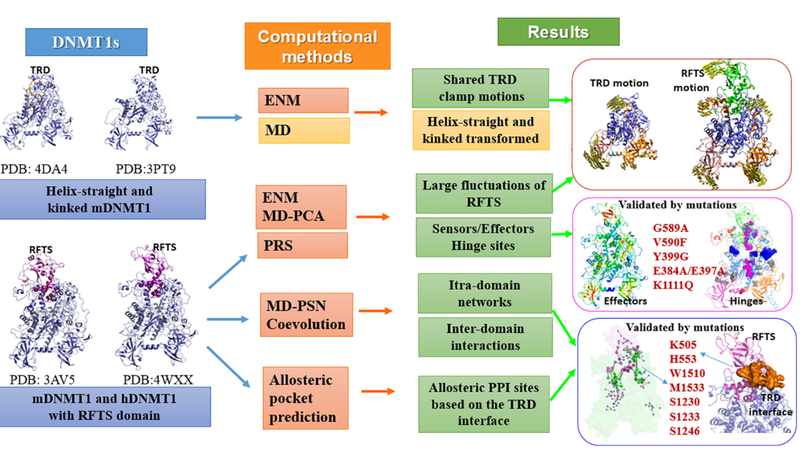
Highlights in the publication include 1)Dynamics and allosteric potentials of the RFTS domain are proposed. 2)Hinge sites located at the RFTS-CD interface are key regulators for inter-domain interactions. 3)Network analysis reveals local allosteric networks and inter-domain communication pathways in DNMT1. 4)A potential allosteric site at the TRD interface for DNMT1 is identified.
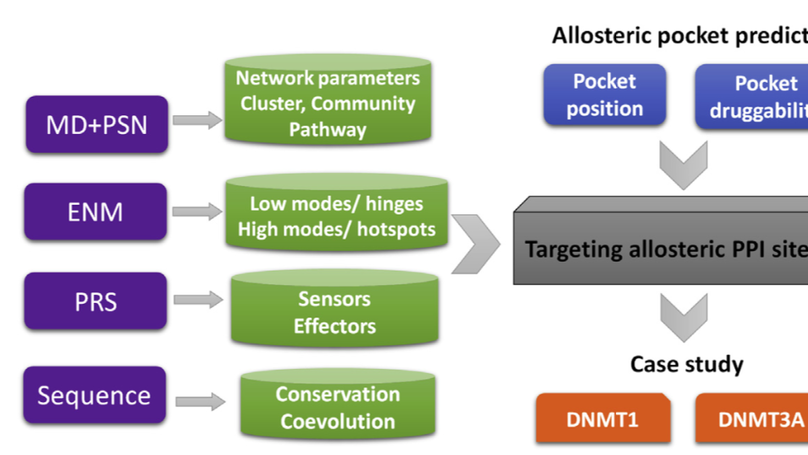
Computational tools now provide reliable assessments of hot spots and pockets found in PPIs, proving reasonable starting points for epigenetic drug design. In this chapter, a hybrid approach, comprising protein structure network, elastic network model, perturbation response scanning and sequence evolution analysis, was proposed for the systematic study of intrinsic dynamics, allostery regulation principle, allosteric sites and pockets. By applied this integrating approach to DNMT1 and DNMT3A, the roles of protein-protein and domain-domain interactions in allosteric drug design were highlighted.
Contact
- yu.zhu.22@ucl.ac.uk
- University College London, London, Bloomsbury WC1E 6BT
- Division of Medicine
- Yu Zhu




